SH3PX1 (SNX9) Human Gene Knockout Kit (CRISPR)
CAT#: KN402822
SNX9 - KN2.0, Human gene knockout kit via CRISPR, non-homology mediated.
KN2.0 knockout kit validation
KN402822 is the updated version of KN202822.
USD 1,290.00
2 Weeks*
Size
Other products for "SNX9"
Specifications
| Product Data | |
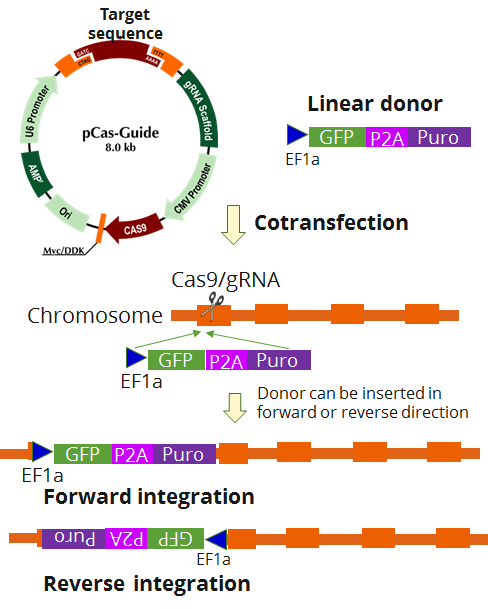
| Format | 2 gRNA vectors, 1 linear donor |
| Donor DNA | EF1a-GFP-P2A-Puro |
| Symbol | SNX9 |
| Locus ID | 51429 |
| Disclaimer | The kit is designed based on the best knowledge of CRISPR technology. The system has been functionally validated for knocking-in the cassette downstream the native promoter. The efficiency of the knock-out varies due to the nature of the biology and the complexity of the experimental process. |
| Reference Data | |
| RefSeq | NM_016224 |
| Synonyms | SDP1; SH3PX1; SH3PXD3A; WISP |
| Summary | This gene encodes a member of the sorting nexin family. Members of this family contain a phosphoinositide binding domain, and are involved in intracellular trafficking. The encoded protein does not contain a coiled coil region, like some family members, but does contain a SRC homology domain near its N-terminus. The encoded protein is reported to have a variety of interaction partners, including of adaptor protein 2 , dynamin, tyrosine kinase non-receptor 2, Wiskott-Aldrich syndrome-like, and ARP3 actin-related protein 3. The encoded protein is implicated in several stages of intracellular trafficking, including endocytosis, macropinocytosis, and F-actin nucleation. [provided by RefSeq, Jul 2013] |
Documents
| Product Manuals |
| FAQs |
| SDS |
Resources
Other Versions
| SKU | Description | Size | Price |
|---|---|---|---|
| GA109800 | SNX9 CRISPRa kit - CRISPR gene activation of human sorting nexin 9 |
USD 1,290.00 |
{0} Product Review(s)
0 Product Review(s)
Submit review
Be the first one to submit a review
Product Citations
*Delivery time may vary from web posted schedule. Occasional delays may occur due to unforeseen
complexities in the preparation of your product. International customers may expect an additional 1-2 weeks
in shipping.






























































































































































































































































 Germany
Germany
 Japan
Japan
 United Kingdom
United Kingdom
 China
China